Precision amplified
Equinox® Library Amplification Kits deliver excellent fidelity, uniform sequence coverage, and high library complexity to specifically address the stringent demands of applications such as rare variant detection, circulating cell-free DNA (cfDNA) analysis, single-cell analysis, and hybridization capture. Kits contain a uniquely engineered, ultra-high fidelity DNA polymerase in an optimized hot start PCR mix formulated for high efficiency, low-bias NGS library amplification.
Key Features and Benefits
- Improve overall assay sensitivity with ultra-high-fidelity amplification, reducing misincorporation events by up to 40%
- Support low-input applications and automated workflows with an effective antibody-based hot start formulation
- Amplify libraries efficiently from a wide range of inputs (0.1 pg to 500 ng) and GC content (15% to 85%)
- Optimize sequencing economy with highly uniform sequence coverage
- Achieve robust performance in hybridization capture workflows due to compatibility with paramagnetic beads
- Leverage the performance benefits of Equinox DNA Polymerase for bisulfite converted DNA with Equinox Uracil Tolerant Library Amplification Kits
Applications
- Low-frequency variant detection NGS assays, including those utilizing challenging samples such as FFPE and cfDNA
- Hybridization capture workflows
- Single-cell analysis
- Whole genome sequencing
- Amplicon sequencing
- RNA-Seq
- ChIP-Seq, ATAC-Seq, and associated epigenetic applications
- Illumina and non-Illumina sample preparation workflows
- Bisulfite-converted DNA
- Damaged DNA samples or templates containing modified bases
Flexible Kit Configurations
Equinox Library Amplification Kits are designed for high-efficiency, high-fidelity amplification of next generation sequencing (NGS) libraries. The ready-to-use mix contains an optimized PCR buffer and hot start enzyme formulation that enables library amplification with minimal bias and error across a broad range of input amounts and GC content, and performance is maintained in the presence of a variety of paramagnetic beads.
Three different Equinox Library Amplification Kits are available, each available with or without amplification primers:
|
The Equinox Library Amplification Kit supports highly sensitive applications with our ultra-high-fidelity polymerase in a convenient 2X master mix format |
|
|
The Equinox HC (High Concentration) Library Amplification Kit provides a more concentrated (4X) master mix, for library amplification reactions from more dilute inputs |
|
|
The Equinox Uracil Tolerant Library Amplification Kit enables amplification of uracil-containing templates including bisulfite-converted, deaminated, or damaged (e.g., FFPE) DNA |
Key Performance Data
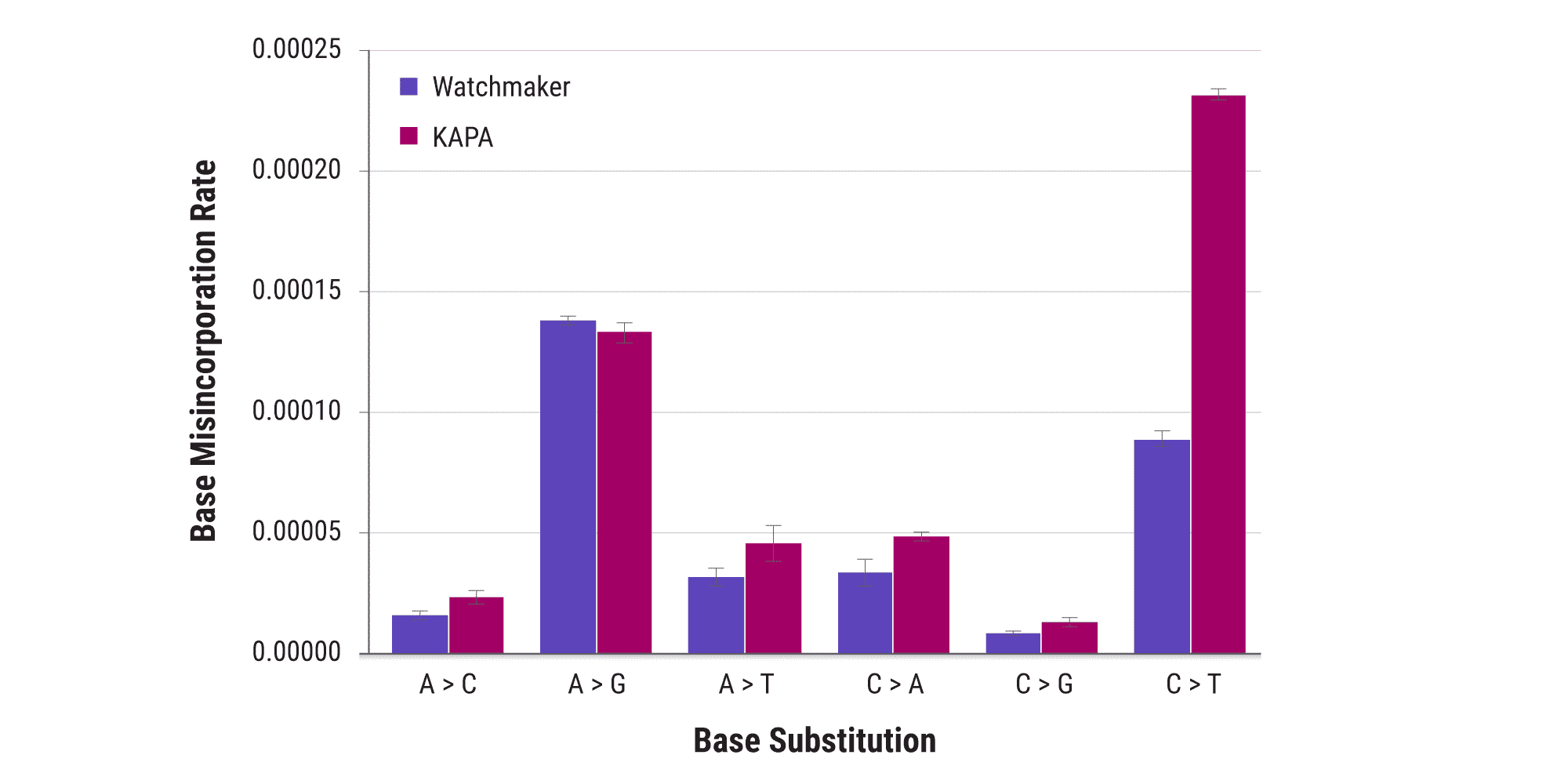
Library amplification with Equinox enables rare mutation detection
Ultra-high-fidelity library amplification is critical for sensitive applications. Equinox is a proprietary proofreading polymerase, capable of a 40% reduction in overall polymerase error rate in comparison to KAPA HiFi HotStart ReadyMix. This enables sensitive variant detection by minimizing overall error rates and reducing false variant calls. A significant reduction in C>T substitutions is particularly important as this mutation is associated with the spontaneous deamination of methylated cytosine to uracil, and is also one of the most common mutation types in cancers.

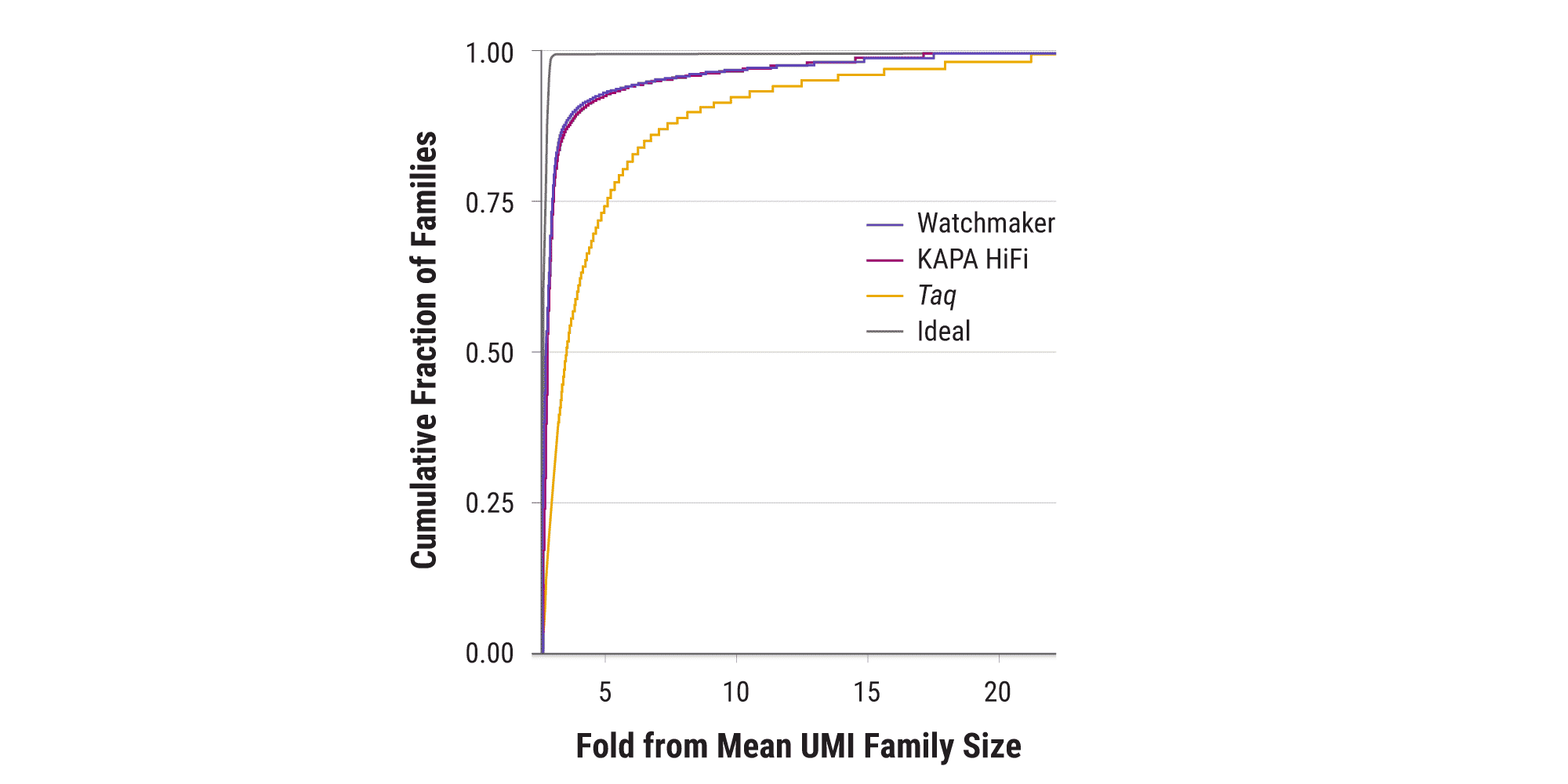
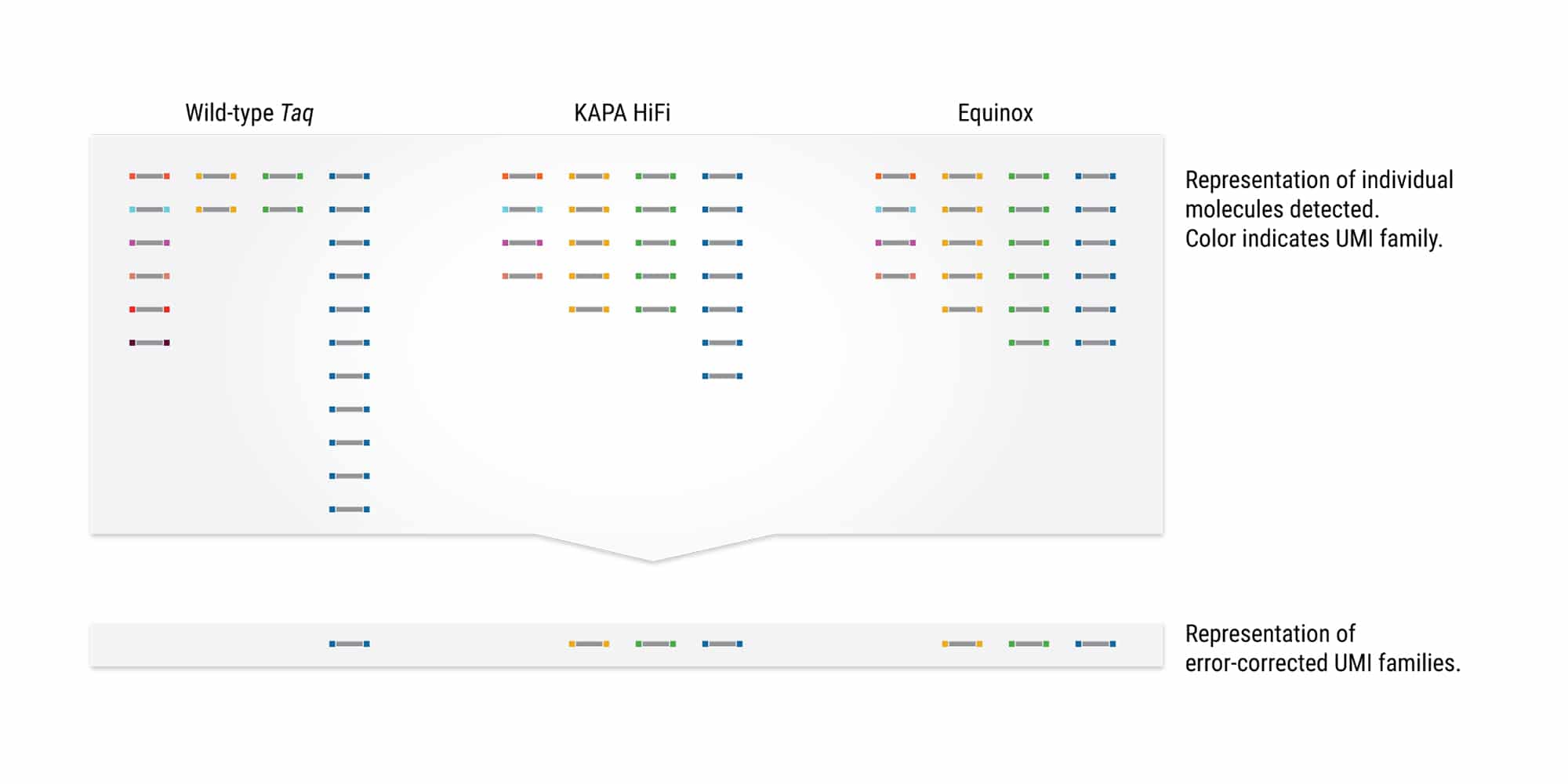
Low-bias amplification delivers uniform UMI family coverage
Unique Molecular Indices (UMIs; also known as molecular barcodes) are added to sequencing libraries prior to PCR amplification to enable accurate bioinformatic identification of PCR duplicates and improve variant calling in low-input applications. Biased amplification—the preferential amplification of a small number of molecules at the expense of others—results in uneven UMI family representation. This generates large numbers of singleton UMIs (families represented by only one read) that cannot be error corrected. A significant amount of additional sequencing may be required to attain requisite coverage of low abundance sequences. Equinox enable uniform UMI family amplification, supporting coverage for >75% of all read families (and >90% of read families with GC content from 25 – 75%) within 3X of the mean family depth.


Effective hot start formulation safeguards low-input applications and automated workflows
The Equinox Amplification Master Mix is formulated with a highly effective hot start antibody that strongly inhibits both the 5’→ 3’ polymerase and 3’ → 5’ exonuclease activities of the enzyme. This mitigates degradation of primers and low-input samples, and minimizes artifacts resulting from nonspecific amplification — thereby improving sensitivity in automated library construction.
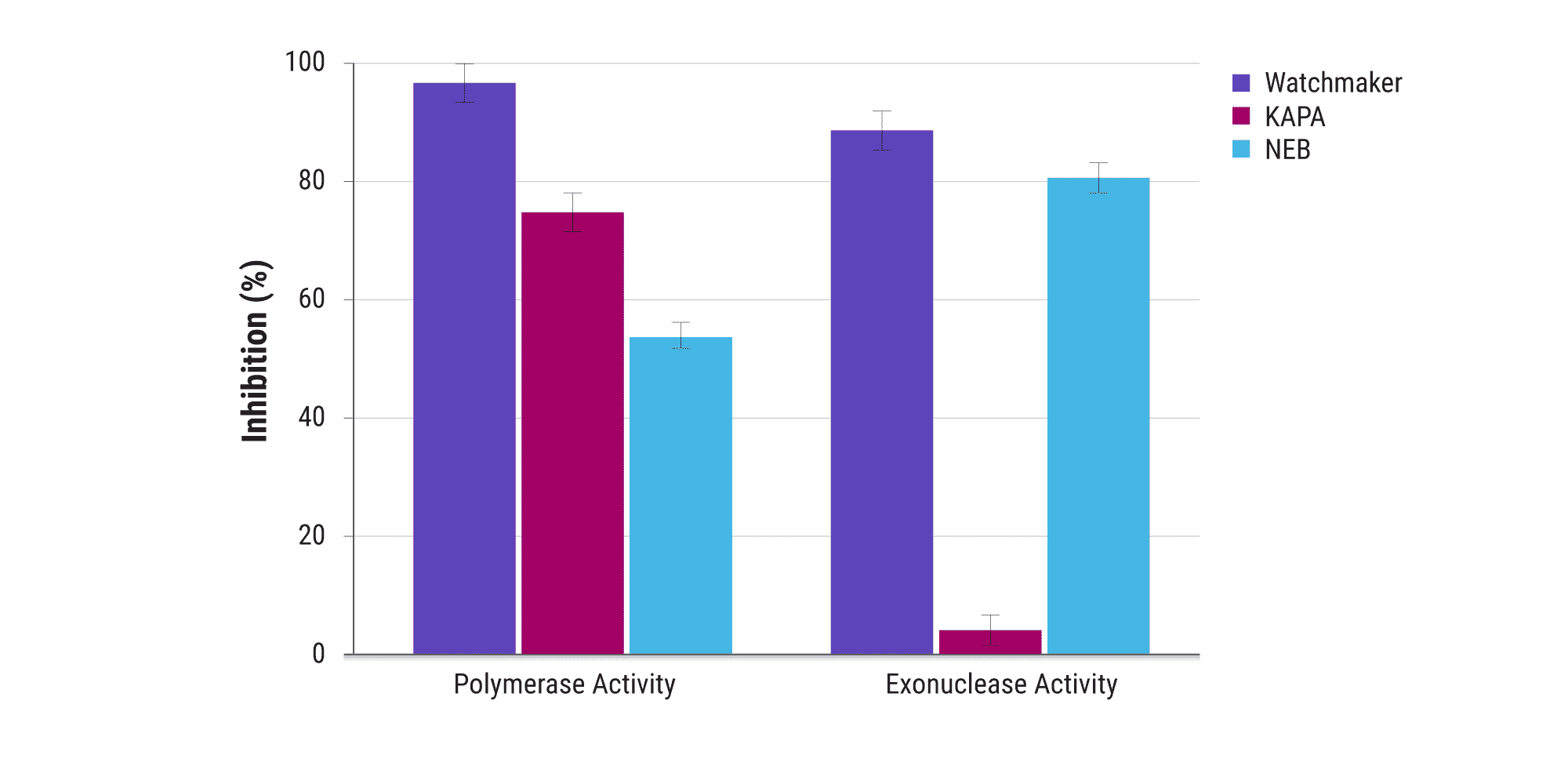

High-efficiency amplification limits bias and artifacts
High efficiency library amplification with Equinox makes it possible to limit the number of cycles needed to generate the yield needed for downstream steps. Amplification efficiency remains robust, even when yields >1 μg are required. This minimizes PCR-associated bias and artifacts for applications such as single-plex hybridization capture.
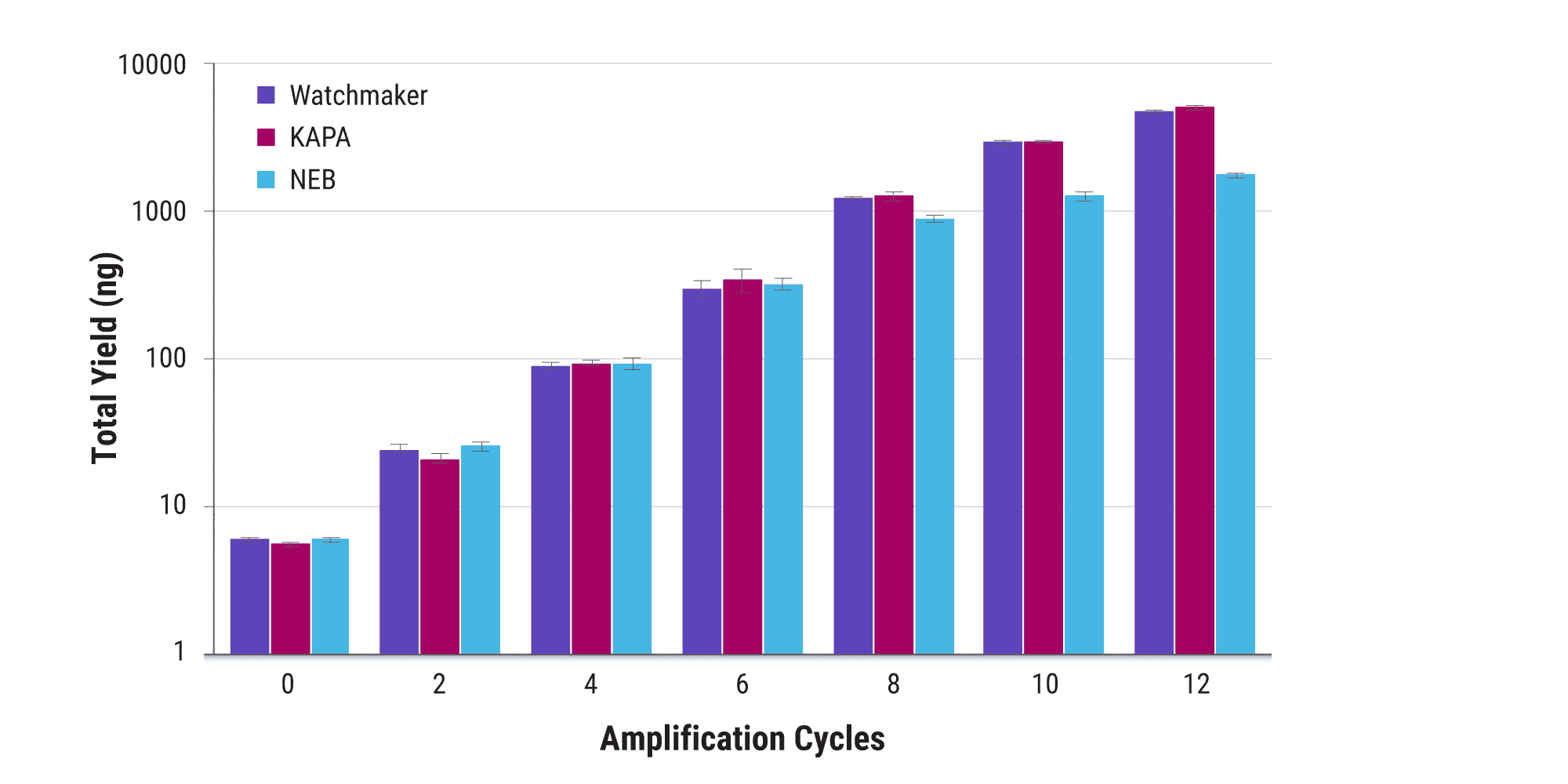

Uniform amplification improves sequencing economy
PCR bias is exacerbated when working with challenging conditions or samples. Equinox introduces minimal amplification bias, even in the presence of paramagnetic beads. This simplifies protocols that benefit from on-bead amplification, such as hybridization capture workflows. Additionally, Equinox delivers even coverage uniformity across complex genomes which can reduce the amount of sequencing needed to achieve desired coverage depths.
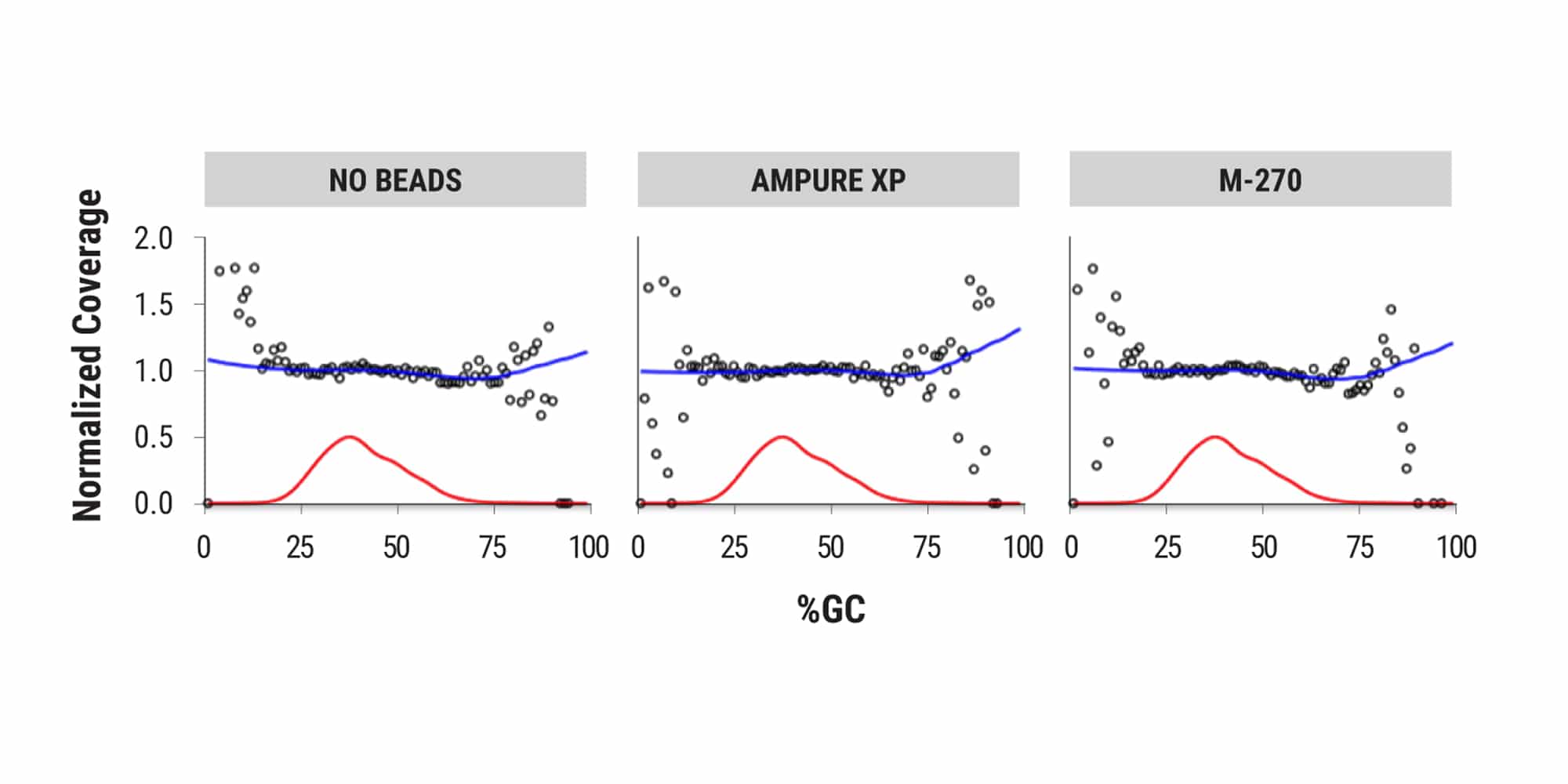
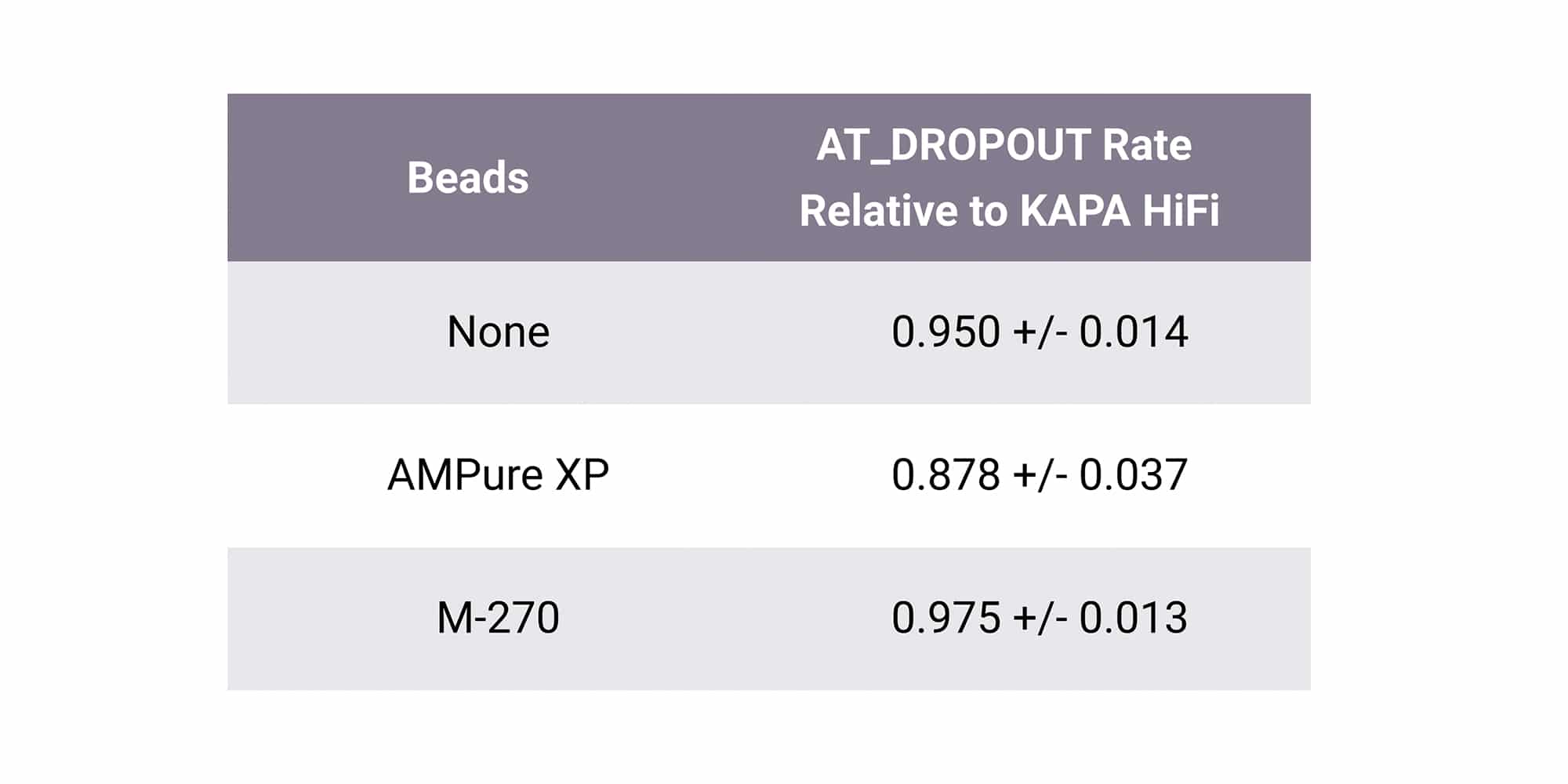


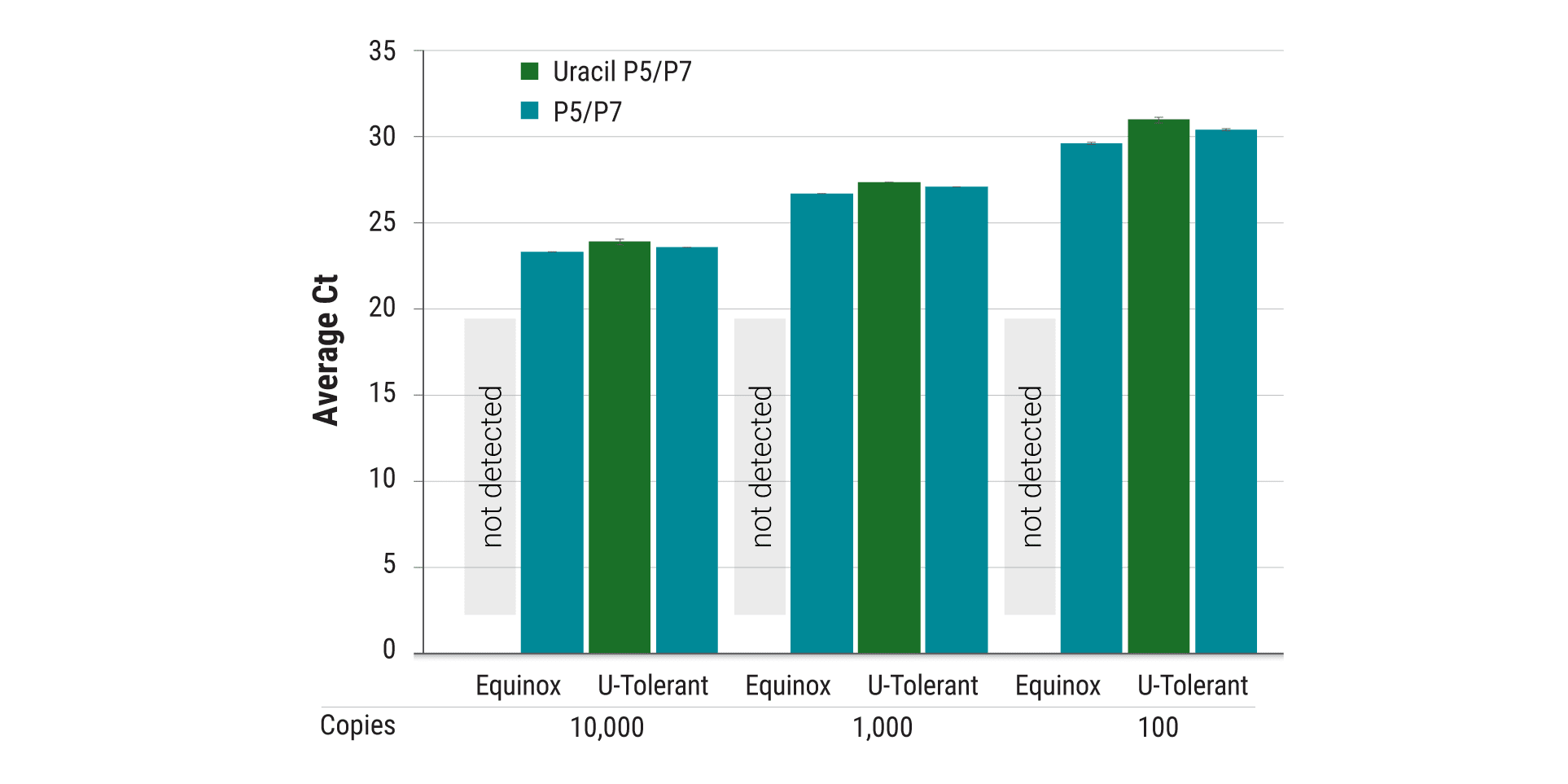
Robust amplification of uracil-containing templates
Many proofreading (B-family) polymerases stall replication in response to uracil or other modified bases in DNA templates. Equinox Uracil Tolerant Polymerase is an engineered polymerase that utilizes uracil-containing templates with high efficiency. This ensures high yields and minimal bias when amplifying bisulfite treated DNA for epigenetic applications.