QUICK LINKS
phi29 Highlights
phi29 DNA Polymerase exhibits strong strand displacement activity and high processivity to enable efficient isothermal DNA amplification from low DNA template amounts. Its 3′ → 5′ exonuclease activity delivers high fidelity amplification, making it an appropriate solution for sequencing DNA template preparation.
- Strong strand displacement activity facilitates isothermal amplification
- Strong processivity delivers products up to 70 kb in length
- High-fidelity amplification supports DNA template preparation for sequencing
- Custom formats available, including high concentration
- Highly stringent enzyme production ensures quality performance across lots
QUICK LINKS
Applications
- Multiple displacement amplification (MDA)
- Rolling circle amplification (RCA)
- Whole genome amplification (WGA)
- Cell-free cloning
- Preparation of DNA template for sequencing
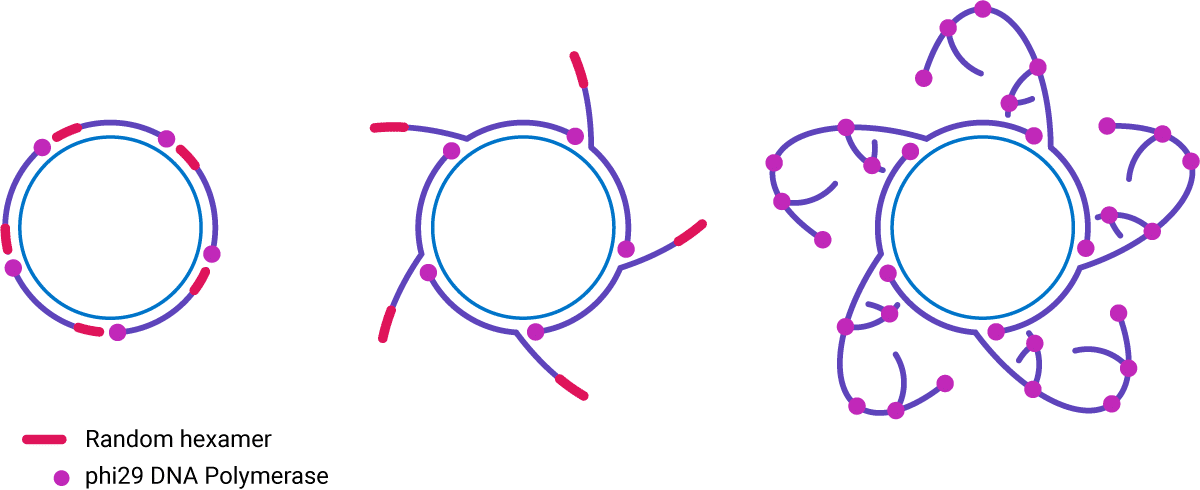
Figure 1. Schematic of rolling circle amplification using phi29 DNA polymerase.
QC Specifications
| Description | Specification |
|---|---|
| Protein Purity Assay | ≥ 99% |
| Exonuclease Assay | Functional |
| Specific Activity | 100,000 – 190,000 U/mg |
| DNA contamination Assay (E. coli, mammalian, library)* | < 10 copies |
| Phosphatase Contamination Assay* | < 1% released |
| Endonuclease Contamination Assay* | Not detectable |
*As assessed using 1,000 U of enzyme input per assay.
Properties
Unit definition: One unit is defined as the amount of enzyme required to convert 50 pmol of dNTPs into a polynucleotide fraction in 10 minutes at 30°C
Reaction conditions:
1X phi29 Pol Reaction Buffer
Incubate at 30°C
Storage Buffer: 10 mM Tris-HCl, pH 7.4, 0.1 mM EDTA, 1 mM DTT, 100 mM KCl, 50% Glycerol, 0.5% Tween
1X phi29 Pol Reaction Buffer: 33 mM Tris-acetate (pH 8.2 at 25°C), 10 mM Mg-acetate, 66 mM K-acetate, 0.1% (v/v) Tween 20, 1 mM DTT
Heat inactivation: 65°C for 10 min
Molecular weight: 66.3 kDa
5’ – 3’ Exonuclease activity: No
3’ – 5’ Exonuclease activity: Yes
Strand Displacement activity: High
Fidelity: High
Processivity: High